NIH P01 CA214091: Noncoding RNAs in Gamma-Herpesvirus Biology and AIDS Malignancies
Co-Investigators |
|
Flemington Lab
|
Project Summary
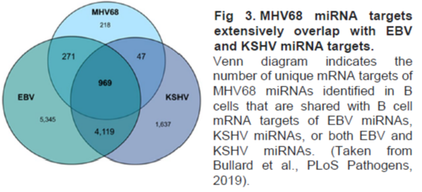
The goal of this program is to use comparative analysis of KSHV, EBV and MHV68 short and long noncoding RNAs to reveal conserved functions of, and regulatory nodes targeted by, γ-herpesvirus noncoding RNAs during tumorigenesis. Our unifying hypothesis is that A) γ-herpesviruses utilize short and long noncoding RNAs to regulate both virus and host gene expression, and B) viral noncoding RNAs and/or virus-perturbed host lncRNAs directly contribute to the genesis of HIV-associated malignancies. In support of this hypothesis, our program has made a number of seminal findings during the first funding period, including: identification of high confidence miRNA targetomes across all three viruses; demonstration of host lncRNA de-regulation in γ-herpesvirus infected cells; discovery of circRNAs across all three viruses; global resolution of EBV and MHV68 transcriptomes and discovery of new noncoding transcripts; demonstration that noncoding RNAs are the predominant viral gene products detectable in some EBV+ malignancies; and demonstration of the first in vivo function for the longstudied EBV EBER1 noncoding RNA. To date, our results from the first cycle have been reported in 20 research publications and 5 review articles.
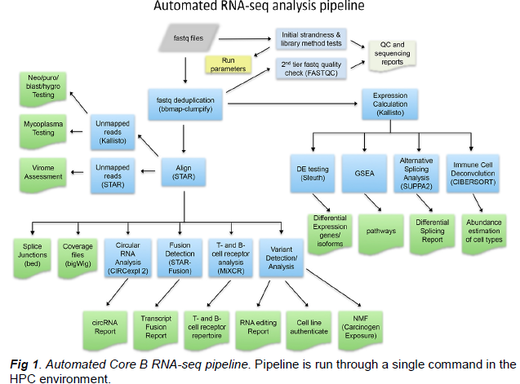
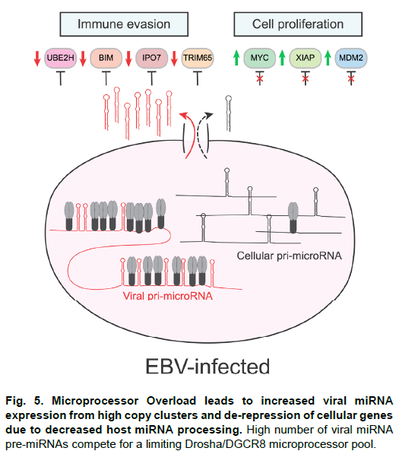
To continue to address our unifying hypothesis we propose three highly integrated projects: Project 1, led by Dr. Renne (University of Florida, UF), will mechanistically study viral lncRNAs and how KSHV-encoded miRNAs induce alterations of host lncRNA expression in the context of HIV-associated KSHV malignancies. Project 2, led by Dr. Flemington (Tulane University) will interrogate the role of EBV (and KSHV) miRNA cluster inhibition of host miRNA maturation through “microprocessor overload” in the context of HIV-associated EBV malignancies. Project 3 led by Dr. Tibbetts (UF) will investigate the function of MHV68 miRNAs and short noncoding RNAs and EBV EBERs in a facile murine chronic infection and tumorigenesis system. The well-organized Administrative core (Core A, Leader: Rolf Renne) will continue to maintain oversight and organization of the program, including biostatistical consultation and adherence to rigor, reproducibility, and transparency standards. Three service cores support all three projects: The RNA-seq and Bioinformatics Core (Core B, Leader: Erik Flemington) at Tulane University will continue to have high impact by developing innovative algorithms and data analyses pipelines. The Recombinant Virus Core (Core C, Leader: Rolf Renne) at UF, which has supported the generation and quality control of more than 60 recombinant viruses, will continue to innovate by implementing CRISPR-Cas-based techniques. The Clinical Sample and Tumorigenesis Core (Core D, Leader: Scott Tibbetts) at UF will support the analysis of a significant numbers of EBV and KSHV-associated tumor samples and perform all proposed tumorigenesis studies. In summary, this program project, which has already had sustained impact on the field, will further increase our understanding of how viral noncoding RNAs and virus-perturbed host lncRNAs contribute to γ-herpesvirus tumorigenesis with the goal of defining new vulnerabilities for therapeutic intervention.
To continue to address our unifying hypothesis we propose three highly integrated projects: Project 1, led by Dr. Renne (University of Florida, UF), will mechanistically study viral lncRNAs and how KSHV-encoded miRNAs induce alterations of host lncRNA expression in the context of HIV-associated KSHV malignancies. Project 2, led by Dr. Flemington (Tulane University) will interrogate the role of EBV (and KSHV) miRNA cluster inhibition of host miRNA maturation through “microprocessor overload” in the context of HIV-associated EBV malignancies. Project 3 led by Dr. Tibbetts (UF) will investigate the function of MHV68 miRNAs and short noncoding RNAs and EBV EBERs in a facile murine chronic infection and tumorigenesis system. The well-organized Administrative core (Core A, Leader: Rolf Renne) will continue to maintain oversight and organization of the program, including biostatistical consultation and adherence to rigor, reproducibility, and transparency standards. Three service cores support all three projects: The RNA-seq and Bioinformatics Core (Core B, Leader: Erik Flemington) at Tulane University will continue to have high impact by developing innovative algorithms and data analyses pipelines. The Recombinant Virus Core (Core C, Leader: Rolf Renne) at UF, which has supported the generation and quality control of more than 60 recombinant viruses, will continue to innovate by implementing CRISPR-Cas-based techniques. The Clinical Sample and Tumorigenesis Core (Core D, Leader: Scott Tibbetts) at UF will support the analysis of a significant numbers of EBV and KSHV-associated tumor samples and perform all proposed tumorigenesis studies. In summary, this program project, which has already had sustained impact on the field, will further increase our understanding of how viral noncoding RNAs and virus-perturbed host lncRNAs contribute to γ-herpesvirus tumorigenesis with the goal of defining new vulnerabilities for therapeutic intervention.
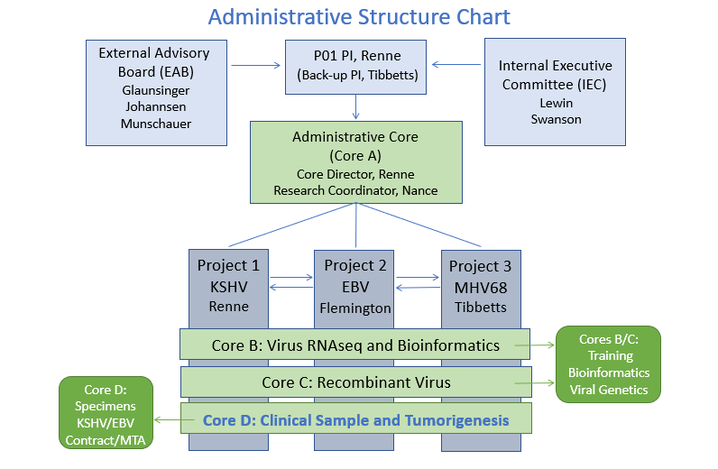
Organizational Chart
Public Health Relevance
The unifying postulate of this competitive renewal P01 proposal is that comparative studies of KSHV, EBV, and MHV68 will continue to accelerate the discovery of pathways regulated by non-coding RNAs (ncRNAs). Our findings over the last 4 years indicate that both short and long ncRNAs contribute to γ-herpesvirus latency and tumorigenesis.
Research Images
External Advisory Board
BRITT GLAUNSINGER, PHD
|
ERIC JOHANNSEN, MD
|
MATHIAS MUNSCHAUER, PHD
|
CHARLES RICE, PHD
|